fMRIPrep: A Robust Preprocessing Pipeline for fMRI Data
fMRIPrep is a NiPreps (NeuroImaging PREProcessing toolS) application (www.nipreps.org) for the preprocessing of task-based and resting-state functional MRI (fMRI).




About
fMRIPrep is a functional magnetic resonance imaging (fMRI) data preprocessing pipeline that is designed to provide an easily accessible, state-of-the-art interface that is robust to variations in scan acquisition protocols and that requires minimal user input, while providing easily interpretable and comprehensive error and output reporting. It performs basic processing steps (coregistration, normalization, unwarping, noise component extraction, segmentation, skull-stripping, etc.) providing outputs that can be easily submitted to a variety of group level analyses, including task-based or resting-state fMRI, graph theory measures, and surface or volume-based statistics.
Note
fMRIPrep performs minimal preprocessing. Here we define ‘minimal preprocessing’ as motion correction, field unwarping, normalization, bias field correction, and brain extraction. See the workflows section of our documentation for more details.
The fMRIPrep pipeline uses a combination of tools from well-known software packages, including FSL, ANTs, FreeSurfer and AFNI. This pipeline was designed to provide the best software implementation for each state of preprocessing, and will be updated as newer and better neuroimaging software become available.
This tool allows you to easily do the following:
Take fMRI data from raw to fully preprocessed form.
Implement tools from different software packages.
Achieve optimal data processing quality by using the best tools available.
Generate preprocessing quality reports, with which the user can easily identify outliers.
Receive verbose output concerning the stage of preprocessing for each subject, including meaningful errors.
Automate and parallelize processing steps, which provides a significant speed-up from manual processing or shell-scripted pipelines.
More information and documentation can be found at https://fmriprep.readthedocs.io/
Principles
fMRIPrep is built around three principles:
Robustness - The pipeline adapts the preprocessing steps depending on the input dataset and should provide results as good as possible independently of scanner make, scanning parameters or presence of additional correction scans (such as fieldmaps).
Ease of use - Thanks to dependence on the BIDS standard, manual parameter input is reduced to a minimum, allowing the pipeline to run in an automatic fashion.
“Glass box” philosophy - Automation should not mean that one should not visually inspect the results or understand the methods. Thus, fMRIPrep provides visual reports for each subject, detailing the accuracy of the most important processing steps. This, combined with the documentation, can help researchers to understand the process and decide which subjects should be kept for the group level analysis.
Citation
Citation boilerplate. Please acknowledge this work using the citation boilerplate that fMRIPrep includes in the visual report generated for every subject processed. For a more detailed description of the citation boilerplate and its relevance, please check out the NiPreps documentation.
Plagiarism disclaimer. The boilerplate text is public domain, distributed under the CC0 license, and we recommend fMRIPrep users to reproduce it verbatim in their works. Therefore, if reviewers and/or editors raise concerns because the text is flagged by automated plagiarism detection, please refer them to the NiPreps community and/or the note to this effect in the boilerplate documentation page.
Papers. fMRIPrep contributors have published two relevant papers: Esteban et al. (2019) [preprint], and Esteban et al. (2020) [preprint].
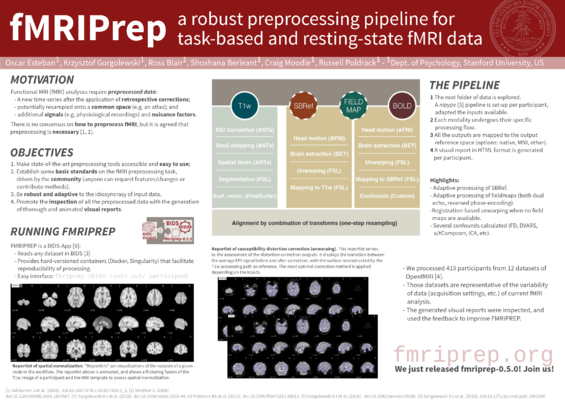
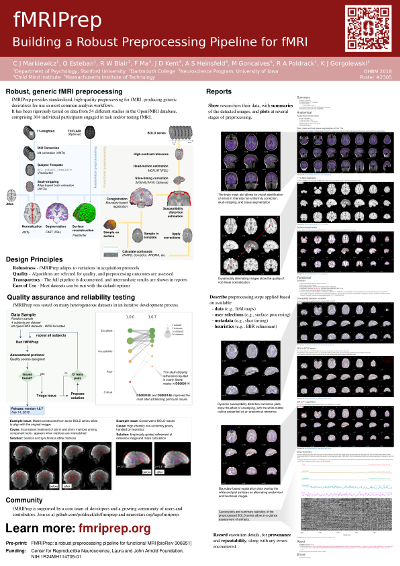
Other. Other materials that have been generated over time include the OHBM 2018 software demonstration and some conference posters:

License information
fMRIPrep adheres to the general licensing guidelines of the NiPreps framework.
License
Copyright (c) 2023, the NiPreps Developers.
As of the 21.0.x pre-release and release series, fMRIPrep is licensed under the Apache License, Version 2.0 (the “License”); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0.
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an “AS IS” BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
Acknowledgements
This work is steered and maintained by the NiPreps Community. This work was supported by the Laura and John Arnold Foundation, the NIH (grant NBIB R01EB020740, PI: Ghosh), and NIMH (R24MH114705, R24MH117179, R01MH121867, PI: Poldrack)
Contents
- Installation
- Usage Notes
- Execution and the BIDS format
- Command-Line Arguments
- Positional Arguments
- Options for filtering BIDS queries
- Options to handle performance
- Options for performing only a subset of the workflow
- Workflow configuration
- Options for modulating outputs
- [DEPRECATED] Options for running ICA_AROMA
- Options relating to confounds
- Specific options for ANTs registrations
- Specific options for handling fieldmaps
- Specific options for SyN distortion correction
- Specific options for FreeSurfer preprocessing
- Options for carbon usage tracking
- Other options
- The command-line interface of the docker wrapper
- Limitations and reasons not to use fMRIPrep
- The FreeSurfer license
- Reusing precomputed derivatives
- Troubleshooting
- About the NiPreps framework licensing
- License information
- Processing pipeline details
- Preprocessing of structural MRI
- BOLD preprocessing
- BOLD reference image estimation
- Head-motion estimation
- Slice time correction
- Susceptibility Distortion Correction (SDC)
- Pre-processed BOLD in native space
- EPI to T1w registration
- Resampling BOLD runs onto standard spaces
- EPI sampled to FreeSurfer surfaces
- HCP Grayordinates
- Confounds estimation
- T2*-driven echo combination
- References
- Outputs of fMRIPrep
- Defining standard and nonstandard spaces where data will be resampled
- FAQ - Frequently Asked Questions
- Should I run quality control of my images before running fMRIPrep?
- What if I find some images have undergone some pre-processing already (e.g., my T1w image is already skull-stripped)?
- My fMRIPrep run is hanging…
- I have already run
recon-allon my subjects, can I reuse my outputs? - ERROR: it appears that
recon-allis already running - Running subjects in parallel
- How much CPU time and RAM should I allocate for a typical fMRIPrep run?
- A new version of fMRIPrep has been published, when should I upgrade?
- I’m running fMRIPrep via Singularity containers - how can I troubleshoot problems?
- What is TemplateFlow for?
- How do you use TemplateFlow in the absence of access to the Internet?
- How do I select only certain files to be input to fMRIPrep?
- Can fMRIPrep continue to run after encountering an error?
- Can I use fMRIPrep for longitudinal studies?
- How to decrease fMRIPrep runtime when working with large datasets?
- Error in slice timing correction: insufficient length of BOLD data after discarding nonsteady-states
- Developers - API
- What’s new
- 23.2.1 (March 06, 2024)
- 23.2.0 (January 10, 2024)
- 23.1.4 (August 1, 2023)
- 23.1.3 (June 24, 2023)
- 23.1.2 (June 16, 2023)
- 23.1.1 (June 14, 2023)
- 23.1.0 (June 12, 2023)
- 23.0.2 (April 24, 2023)
- 23.0.1 (March 24, 2023)
- 23.0.0 (March 13, 2023)
- 22.1.1 (January 04, 2023)
- 22.1.0 (December 12, 2022)
- 22.0.2 (September 27, 2022)
- 22.0.1 (September 13, 2022)
- 22.0.0 (July 28, 2022)
- 21.0.4 (September 29, 2022)
- 21.0.3 (September 6, 2022)
- 21.0.2 (April 21, 2022)
- 21.0.1 (January 24, 2022)
- 21.0.0 (December 14, 2021)
- 20.2.7 (January 24, 2022)
- 20.2.6 (October 27, 2021)
- 20.2.5 (October 12, 2021)
- 20.2.4 (October 04, 2021)
- 20.2.3 (July 21, 2021)
- 20.2.2 (July 16, 2021)
- 20.2.1 (November 06, 2020)
- 20.2.0 (September 28, 2020)
- 20.1.4 (July 16, 2021)
- 20.1.3 (September 15, 2020)
- 20.1.2 (September 04, 2020)
- 20.1.1 (June 04, 2020)
- 20.1.0 (May 27, 2020)
- 20.0.x series (February 2020)
- 1.5.x series (September 2019)
- 1.4.x series (May 2019)
- 1.3.x series (March 2019)
- 1.2.x series (January 2019)
- 1.1.x series (October 2018)
- 1.0.x series (May 2018)
- 1.0.15 (May 17, 2018)
- 1.0.14 (May 15, 2018)
- 1.0.13 (May 11, 2018)
- 1.0.12 (May 03, 2018)
- 1.0.11 (April 16, 2018)
- 1.0.10 (April 16, 2018)
- 1.0.9 (April 10, 2018)
- 1.0.8 (February 22, 2018)
- 1.0.7 (February 13, 2018)
- 1.0.6 (29th of January 2018)
- 1.0.5 (21st of January 2018)
- 1.0.4 (15th of January 2018)
- 1.0.3 (3rd of January 2018)
- 1.0.2 (2nd of January 2018)
- 1.0.1 (1st of January 2018)
- 1.0.0 (6th of December 2017)
- 1.0.0-rc13 (1st of December 2017)
- 1.0.0-rc12 (29th of November 2017)
- 1.0.0-rc11 (24th of November 2017)
- 1.0.0-rc10 (9th of November 2017)
- 1.0.0-rc9 (2nd of November 2017)
- 1.0.0-rc8 (27th of October 2017)
- 1.0.0-rc7 (20th of October 2017)
- 1.0.0-rc6 (11th of October 2017)
- 1.0.0-rc5 (25th of September 2017)
- 1.0.0-rc4 (12th of September 2017)
- 1.0.0-rc3 (28th of August 2017)
- 1.0.0-rc2 (12th of August 2017)
- 1.0.0-rc1 (8th of August 2017)
- 0.x series (July 2017)
- 0.6.0 (31st of July 2017)
- 0.5.4 (20th of July 2017)
- 0.5.3 (18th of July 2017)
- 0.5.2 (30th of June 2017)
- 0.5.1 (24th of June 2017)
- 0.5.0 (21st of June 2017)
- 0.4.6 (14th of June 2017)
- 0.4.5 (12th of June 2017)
- 0.4.4 (20th of May 2017)
- 0.4.3 (10th of May 2017)
- 0.4.2 (3rd of May 2017)
- 0.4.1 (20th of April 2017)
- 0.4.0 (20th of April 2017)
- 0.3.2 (7th of April 2017)
- 0.3.1 (24th of March 2017)
- 0.3.0 (20th of March 2017)
- 0.2.0 (13th of January 2017)
- 0.1.2 (3rd of October 2016)
- 0.1.1 (30th of July 2016)
- 0.0.1